Everyone Is Building AI Agents for Drug Discovery. We Built a Workflow for One Task. Here's Why.
Why we rebuilt CyteType as a deterministic AI workflow instead of an agent, and why that distinction matters for production cell annotation pipelines.
The dominant narrative in life sciences AI right now is autonomous agents. A Stanford spinout raised $13.5M to build a platform with 300+ scientific tools. A publicly traded company describes its system as "pharmaceutical superintelligence," running on LangChain and LangGraph with 140+ skills and over 1,000 tools. A $70M startup claims to compress six months of analysis into 24 hours. A London-based startup offers an AI bioinformatician at a third of the time and cost.
Every one of these positions agents as the architecture for scientific automation. We went the other direction.
In January 2026, we rebuilt CyteType from a multi-agent system into a deterministic AI workflow: one task, one workflow, no autonomous planning loops. If you want to understand what broke in production with the previous architecture, our previous post covers that story in full. This post is about the broader argument: when agents are the right choice, when they aren't, and how to think about the distinction as a computational biology team.
What an agent is, precisely
The term gets used loosely enough to be almost meaningless, so it's worth being precise. An agent takes a natural language goal, plans the steps it believes will achieve that goal, executes tools or API calls, observes the results, and iterates until it decides the goal is met. Planning is the defining feature. The model doesn't just respond to a query; it decides what to do next at each step.
Workflows work differently. The developer defines the sequence in advance. LLMs are used within that sequence to handle the parts that require natural language reasoning, but the program controls when each step runs and what happens between them. The model is transactional; the structure is deterministic.
The practical difference is who controls depth and breadth: the model, or the developer.
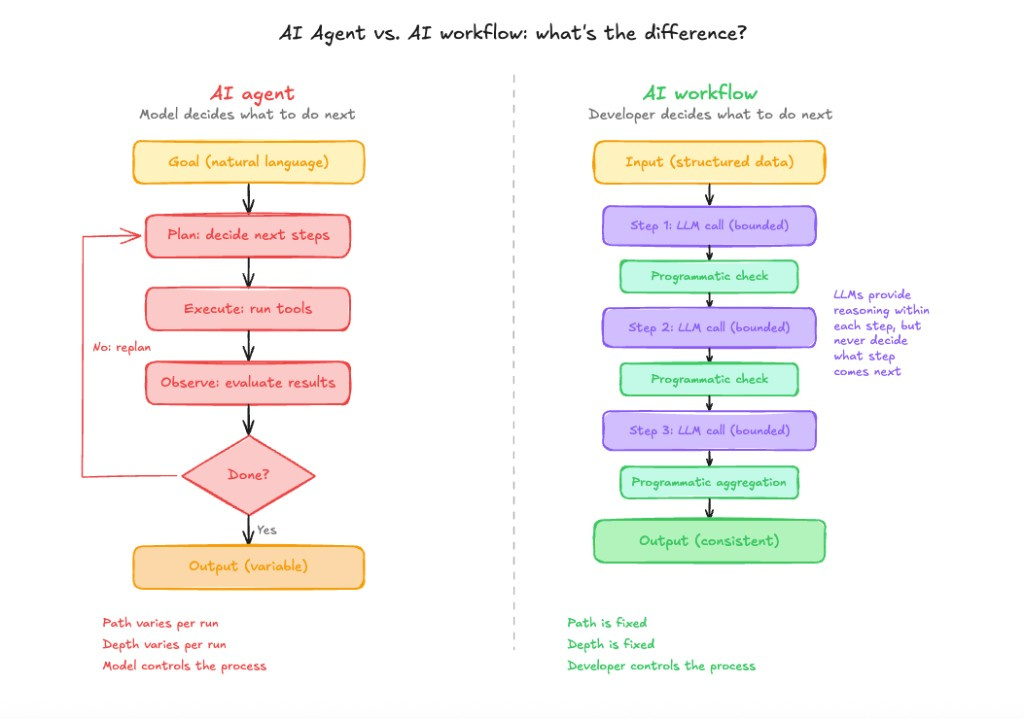
Agent architectures follow a plan-execute-observe pattern; the model decides what to do next at each iteration. Deterministic AI workflows interleave bounded LLM calls with programmatic checks in a fixed order. The key difference is who controls depth and breadth: the model or the developer. In the diagram, purple marks LLM steps and green marks programmatic steps.
The trade-off nobody talks about
A general-purpose agent with 300 tools is necessarily shallow on each of them. The planning loop distributes attention across a large tool surface, and with that many options, attention is diffuse. For tasks that require deep, contextual reasoning within a narrow domain, that shallowness is a real problem.
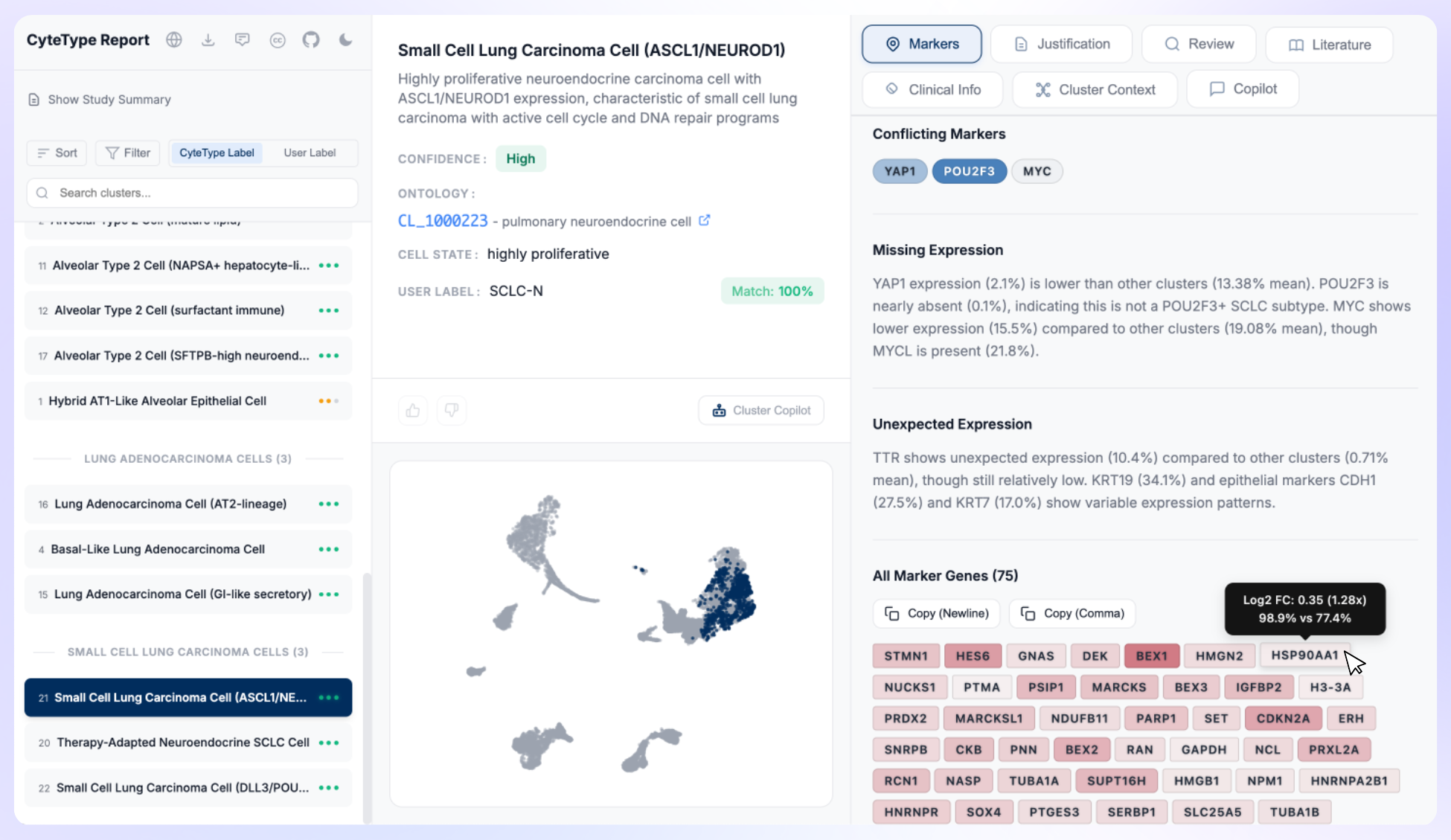
Cell type annotation is a useful example. It's not a single lookup or a straightforward classification. Annotating a cluster correctly requires reasoning across tissue-specific marker expression patterns, the relationship between co-expressed genes, the disease state of the sample, species context, and often the relationship between similar populations that differ in subtle but biologically meaningful ways. Annotating CD8+ T cell subtypes in a tumour-infiltrating lymphocyte context is not the same problem as annotating the same markers in a healthy spleen, and the biological significance of the distinction is real.
Getting this right requires depth. Our constraint of doing one thing is what makes that depth possible.
Where agents genuinely work
There are tasks where agent architectures make real sense. The common factor is that the task is exploratory, one-off, and the acceptable variance in outputs is relatively high.
A biologist investigating a new dataset for the first time, forming hypotheses about which populations are present, or designing follow-up experiments is doing exactly this kind of work. The goal is open-ended, the tool use is genuinely varied, and the output doesn't need to be reproducible across runs. Speed of exploration matters more than consistency.
Several platforms are producing meaningful results in this space. One has demonstrated compression of transcriptomic analyses from weeks to hours in exploratory contexts. Another has reported validated pathway-level discoveries in cardiac fibrosis and neurodegeneration. These are real contributions. Agent architectures fit this problem well because the task is inherently exploratory, and the researcher remains in the loop to evaluate outputs as they emerge.
Where agents break down
The failure mode is predictable, and it's not an implementation problem: it's an architectural mismatch.
Cell type annotation is a production task for most pharma organisations. It runs across every single-cell dataset the team generates. The annotations feed directly into downstream decisions: which populations to target, how to stratify patient cohorts, which findings to carry forward into validation experiments. Getting different answers on the same dataset across two separate runs isn't a minor inconvenience. It's a reproducibility problem with concrete downstream consequences.
The specific failure modes we documented in our own multi-agent system, and subsequently in others, are: run-to-run variance in annotations for identical inputs; selective evidence gathering, where the model finds evidence supporting an early hypothesis and stops looking; and inconsistent analytical depth across clusters, where straightforward clusters receive thorough treatment while ambiguous ones receive plausible-sounding but poorly-supported annotations.
These aren't implementation bugs. They're emergent properties of goal-directed architectures. A system optimised to produce an answer will generate confident annotations on genuinely ambiguous clusters because the architecture provides no mechanism to surface "evidence is insufficient." That's a problem in any context. In a production pipeline feeding target selection, it's a serious one.
Matching architecture to task type
The practical question isn't "agents or workflows?" It's "which architecture fits this specific task?"
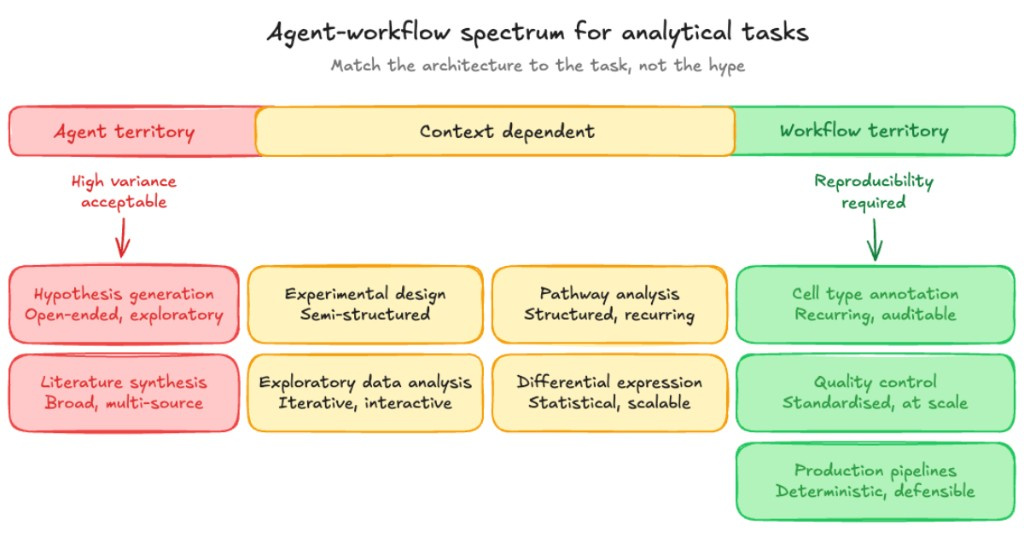
The bar runs from settings where high variance is acceptable through context-dependent work to workflow territory where reproducibility is required. Placement depends on the task: exploratory work clusters on the left; standardised, auditable pipeline steps on the right; several analyses sit in between when the right architecture depends on how you run the study.
The key question is whether the output needs to be defensible, auditable, and consistent across runs. If yes, use a workflow. Control the depth, breadth, and reasoning structure programmatically, not through the model's planning decisions. If no, an agent can let the model explore freely, with the researcher there to evaluate outputs.
A mature drug discovery computational biology team needs both. Agents for exploration, hypothesis generation, and literature synthesis; workflows for annotation, quality control, differential expression, and anything that feeds a downstream decision. The mistake is assuming agent architectures can cover both reliably at scale.
What the results show
CyteType's deterministic AI workflow was validated across 977 clusters in 20 independent datasets. Against comparable LLM-based annotation tools, the benchmarks from our preprint are: 388% improvement over GPTCellType, 268% over CellTypist, and 101% over SingleR.
The largest performance gains came on the hardest cases: ambiguous cell states, rare populations, and disease-perturbed tissues where reference data is limited or absent. These are precisely the populations where annotation quality matters most for drug discovery applications, and where shallow or inconsistent methods tend to fail silently. The workflow also flags when evidence is genuinely insufficient rather than producing a confident-sounding annotation that obscures the uncertainty.
Evaluating tools for your pipeline
If you're assessing AI-based annotation for a production pipeline, three tests are worth running before committing.
For agent-based tools: run the same dataset three times and compare annotations across runs. If the variance is unacceptable for your use case, the architecture is a mismatch for production use, regardless of how good the average annotation looks.
For workflow-based tools: ask what happens when a cluster can't be confidently annotated. If the system always produces an annotation regardless of evidence quality, investigate how uncertainty is handled internally. A tool that suppresses uncertainty generates confident-sounding errors, which are harder to catch downstream than obvious failures.
For your own pipeline: map your computational biology tasks on the spectrum from exploratory to production-grade. Annotation, QC, and differential expression belong in the workflow category. Hypothesis generation, literature synthesis, and experimental design are well-suited to agent-based tools. Treat the architecture decision as a property of the task, not a blanket choice for the whole computational stack.
CyteType is available via pip install cytetype (Python) and from GitHub (R via CyteTypeR). If you're evaluating AI-based annotation tools for your single-cell pipeline, we'd welcome the conversation. Schedule a demo →