Integrating Multi-Omics Data for Effective Target Identification in Drug Discovery
Discover how multi-omics integration is reshaping drug discovery by uncovering disease mechanisms, prioritizing drug targets, and connecting genomics, epigenomics, transcriptomics, proteomics, and metabolomics into a usable biological model.
Uncovering Complex Biology with Multi-Omics
Drug discovery fails when target biology is thinner than it looks. A gene can appear differentially expressed without being causally relevant. A variant can look compelling in genetics but act through a cell state or pathway that is not obvious from bulk measurements alone. A protein can be abundant but not active in the cellular context that matters. Multi-omics helps reduce that mismatch by connecting molecular layers instead of treating each layer as a separate answer.
That matters most in early discovery, where the real task is not to generate more candidate targets. It is to rank targets by mechanistic plausibility, cellular specificity, tractability, and translational relevance.
The Role of Multi-Omics in Early-Stage Drug Discovery Pipelines
In practice, multi-omics is useful when it changes the confidence of a decision. The strongest use cases are not "we collected many omics layers," but questions like these:
- Does human genetics support the target?
- In which cell type or disease state is the target active?
- Is the signal transcriptional, epigenetic, post-transcriptional, or protein-level?
- Does pathway activity converge across modalities?
- Is there evidence that perturbing the target shifts the disease program in the expected direction?
A good integrated analysis can help at three levels:
- Target prioritization
- Combine genetic association, expression specificity, chromatin accessibility, protein abundance, and pathway context.
- Filter out targets that are statistically interesting but biologically weak.
-
Rank candidates by consistency across orthogonal evidence.
-
Patient and disease stratification
- Resolve subtypes that look similar histologically but differ molecularly.
- Identify response-associated programs, resistant states, or lineage plasticity.
-
Define populations where a mechanism is enriched enough to matter clinically.
-
Mechanistic validation
- Connect regulatory programs to downstream transcriptional and phenotypic effects.
- Test whether the proposed mechanism survives cross-modal scrutiny.
- Highlight where follow-up experiments should go first.
What Multi-Omics Actually Adds Beyond Single-Omics
Each layer answers a different biological question.
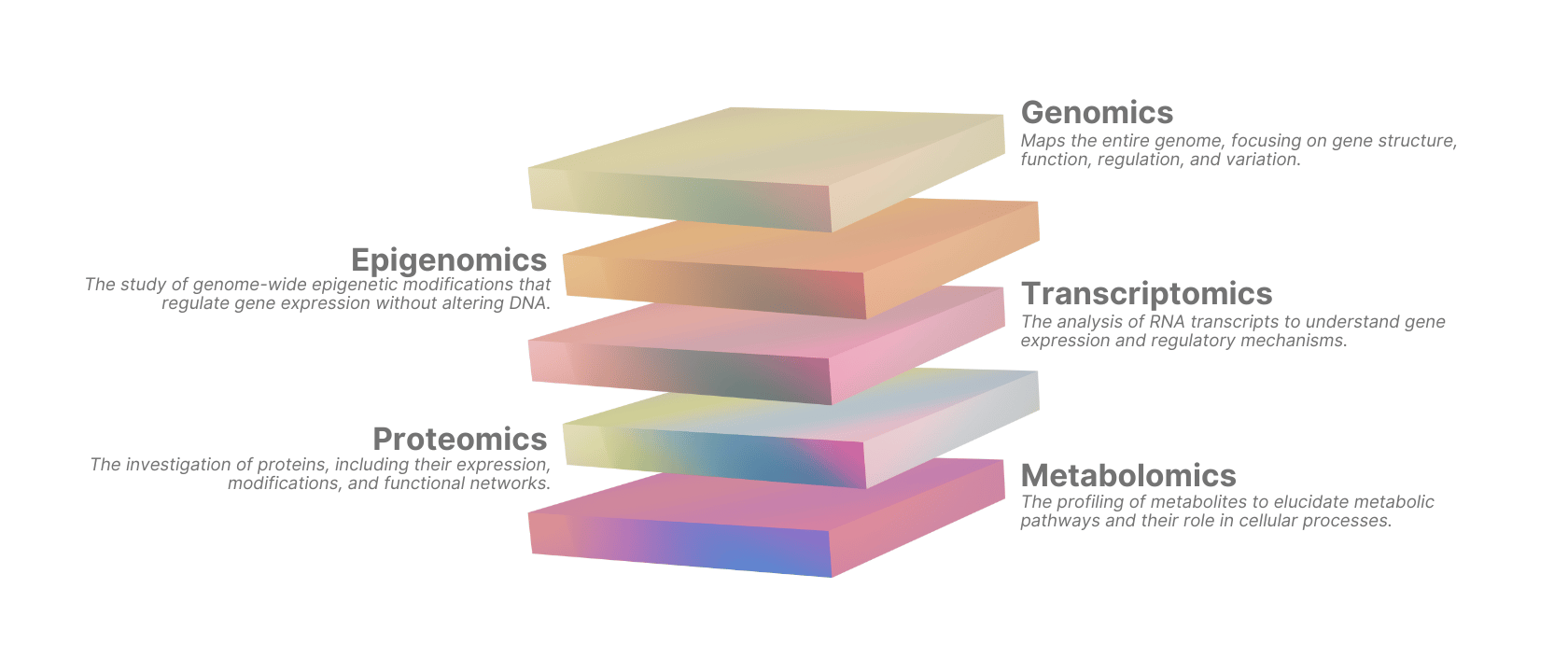
- Genomics helps with inherited risk, somatic alterations, and target support from human biology.
- Epigenomics helps explain which regulatory programs are active and in which cellular context.
- Transcriptomics captures state changes, pathway activity, and heterogeneity.
- Proteomics gets closer to functional effectors, especially where RNA and protein do not align.
- Metabolomics can reveal pathway flux constraints and phenotype-linked metabolic dependencies.
The value comes from convergence. If a target sits near a disease-associated locus, shows cell-type-specific accessibility, drives a transcriptional program in the relevant compartment, and is supported by protein or perturbation readouts, it is harder to dismiss as noise.
Challenges in Multi-Omics Data Integration
This is where many analyses become less useful than they look on slides.
1. Data are rarely matched cleanly
Not every study profiles every modality in the same sample, let alone the same cell. Cohort composition, sample handling, chemistry, and preprocessing can introduce stronger structure than the biology you care about.
2. Modalities do not behave on the same scale
RNA counts, chromatin peaks, protein measurements, and metabolite intensities have different noise models, sparsity profiles, and dynamic ranges. Treating them as interchangeable matrices usually produces brittle results.
3. Integration can over-correct or over-simplify
A joint latent space is useful, but it is not automatically truthful. If the model removes biologically real condition effects because they resemble batch structure, target ranking can become cleaner and less correct at the same time.
4. Interpretation is still the bottleneck
Many teams can generate an embedding. Fewer can explain which cross-modal signal actually strengthens a target hypothesis, what is shared versus modality-specific, and which claims are robust enough for experimental follow-up.
Advancements in Multi-Omics Integration Techniques
The method should match the design of the study.
Unsupervised latent factor models
For cohort-level integration across several molecular layers, factor models remain useful because they separate shared and modality-specific axes of variation. Methods such as MOFA+ are still relevant when the goal is to identify interpretable factors tied to disease state, tissue context, or treatment response.
Probabilistic models for single-cell multimodal data
For paired single-cell measurements, the field has moved toward probabilistic models that explicitly account for technical noise, missing modalities, and batch structure. totalVI is widely used for CITE-seq because it models RNA and protein jointly, including protein background. MultiVI extends this idea to paired or partially paired RNA and chromatin accessibility data, which is especially useful in regulatory target discovery.
Network and pathway-centered integration
When the practical question is target nomination rather than representation learning, network-based integration is often easier to act on. Regulatory networks, protein interaction maps, ligand-receptor frameworks, and pathway models can convert multi-layer evidence into something more decision-friendly.
Benchmarking has improved, but there is no universal winner
Recent benchmarking work in multimodal single-cell integration has been useful because it makes the trade-offs clearer. Methods differ depending on whether the task is modality matching, batch removal, cell-type conservation, imputation, or downstream biological recovery. The best method is task-dependent.
Single-Cell Multi-Omics in Drug Discovery
This is where the biology gets more concrete.
Single-cell multi-omics is particularly useful when disease signal is state-dependent, rare, or compartment-specific. In oncology, immunology, fibrosis, and neurodegeneration, the key mechanism often sits in a minority population, a transient transition state, or a cell-cell interaction that bulk assays dilute away.
Examples where single-cell multi-omics can materially improve discovery work:
- Targeting pathogenic cell states
- Link accessible regulatory regions to the transcriptional programs that define a pathogenic state.
-
Distinguish stable lineages from transient activation states.
-
Resolving mechanism in the microenvironment
- Use transcriptomic, surface protein, and spatial context to identify signaling relationships between compartments.
-
Avoid choosing targets that look compelling in aggregate but are not accessible or relevant in the active niche.
-
Understanding resistance and relapse
- Detect state transitions associated with drug tolerance.
-
Separate pre-existing resistant populations from treatment-induced remodeling.
-
Improving translational alignment
- Map preclinical models against patient-derived cellular states.
- Check whether the biology a drug modulates in vitro is the biology that exists in human tissue.
Applications of Multi-Omics in Target Discovery
Pathway analysis and regulatory modeling
The most useful target programs are often not single genes but network positions. Multi-omics helps identify whether a candidate sits upstream, downstream, or parallel to the disease program, and whether its effect is likely to be cell-intrinsic or microenvironment-mediated.
Biomarker discovery and patient selection
Integrated signatures can support biomarker strategies that are mechanistically tied to response, rather than just correlated with it. This is valuable both for enrichment and for understanding likely non-responders early.
Functional follow-up
Multi-omics is strongest when paired with perturbation. CRISPR screens, Perturb-seq, functional proteomics, or targeted validation assays are what turn target hypotheses into a development decision.
Data Integration Strategies in Early-Stage Drug Pipelines
A useful operating model is to think in three layers:
- Evidence assembly
- Bring together genetics, expression, accessibility, protein, phenotype, and prior knowledge.
-
Normalize identifiers, metadata, and sample structure before attempting joint analysis.
-
Biological integration
- Choose methods based on the question: latent factors for shared structure, probabilistic models for paired single-cell data, network models for mechanistic ranking.
-
Keep batch effects, donor structure, and modality-specific noise explicit.
-
Decision framing
- Rank targets by convergent evidence, not by one modality alone.
- Separate exploratory signal from evidence strong enough to guide spend.
- Push the shortlist into orthogonal validation quickly.
Multi-Omics Data Is Changing How Targets Are Chosen
Multi-omics does not replace experimental biology. It makes early target selection less blind.
The real gain is not that every program suddenly becomes straightforward. The gain is that weak targets are easier to deprioritize, cell-state-specific mechanisms are easier to see, and translational questions can be asked earlier, before cost accumulates around the wrong hypothesis.
For teams working in single-cell and multimodal data, the next step is not collecting more layers for the sake of it. It is building analyses that connect those layers to a concrete target decision.
Multi-Omics Integration with Nygen Analytics
Nygen helps research teams work through complex single-cell and multi-omics datasets without turning every analysis step into a separate engineering project. That matters when the goal is to move from data to biological interpretation quickly, while keeping enough structure and traceability for downstream review.
Where this becomes useful in practice:
- comparing disease and control programs across samples or cohorts
- exploring cell-state heterogeneity in target-relevant compartments
- visualizing integrated readouts in a format experimental teams can work with
- shortening the path from raw data to a biologically interpretable shortlist
If your work depends on identifying target-relevant states, comparing cohorts, or making sense of complex multimodal datasets, explore how Nygen can support that analysis workflow.



