
Public single-cell RNA-seq databases worth using in 2026
An updated guide to the most useful public single-cell RNA-seq databases in 2026, including archives, atlas portals, and domain-specific resources for data discovery and reuse.
Read more →A curated index of articles, updates, events, and published work across single-cell research.

An updated guide to the most useful public single-cell RNA-seq databases in 2026, including archives, atlas portals, and domain-specific resources for data discovery and reuse.
Read more →
Uncover the top single-cell RNA sequencing (scRNA-seq) and multi-omics data analytics tools of 2025. This comprehensive guide reviews 8 platforms to facilitate data analysis and reveal cellular insights.
Read more →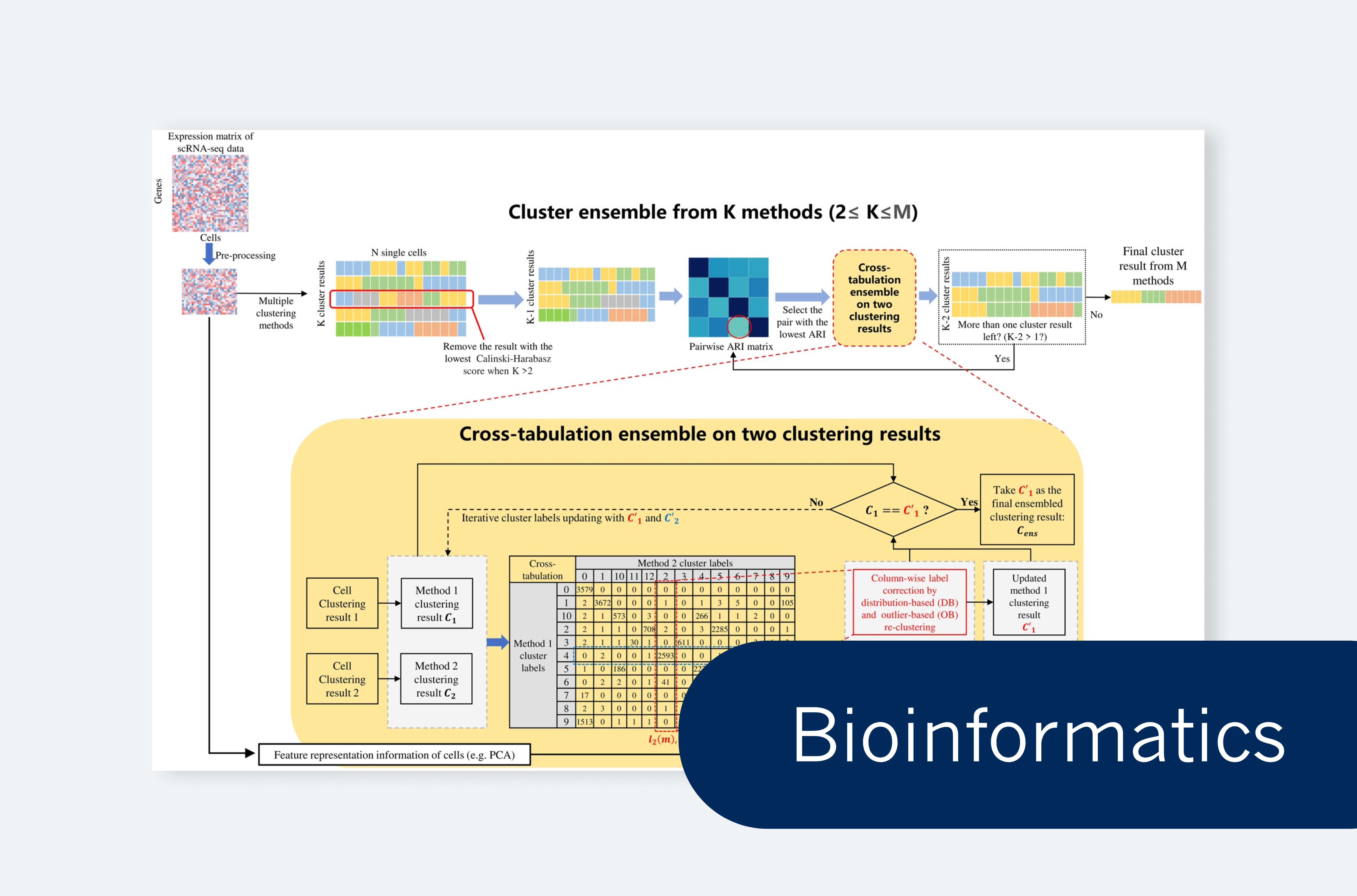
Liang Wang et al. present CTEC, an ensemble clustering method for robust scRNA-seq data analysis.
Read paper →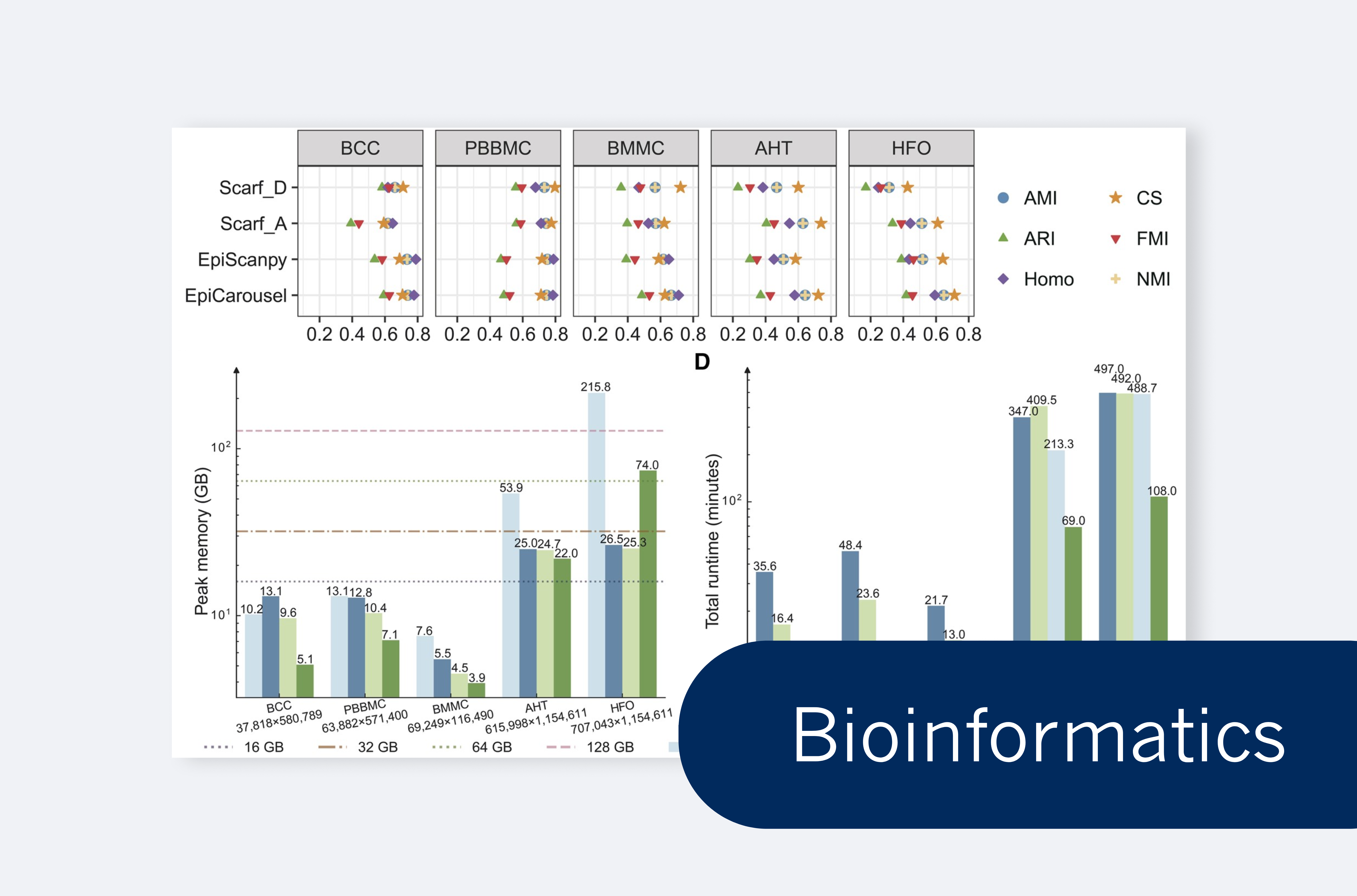
Sijie Li et al. introduce EpiCarousel for efficient metacell identification in large-scale chromatin accessibility datasets.
Read paper →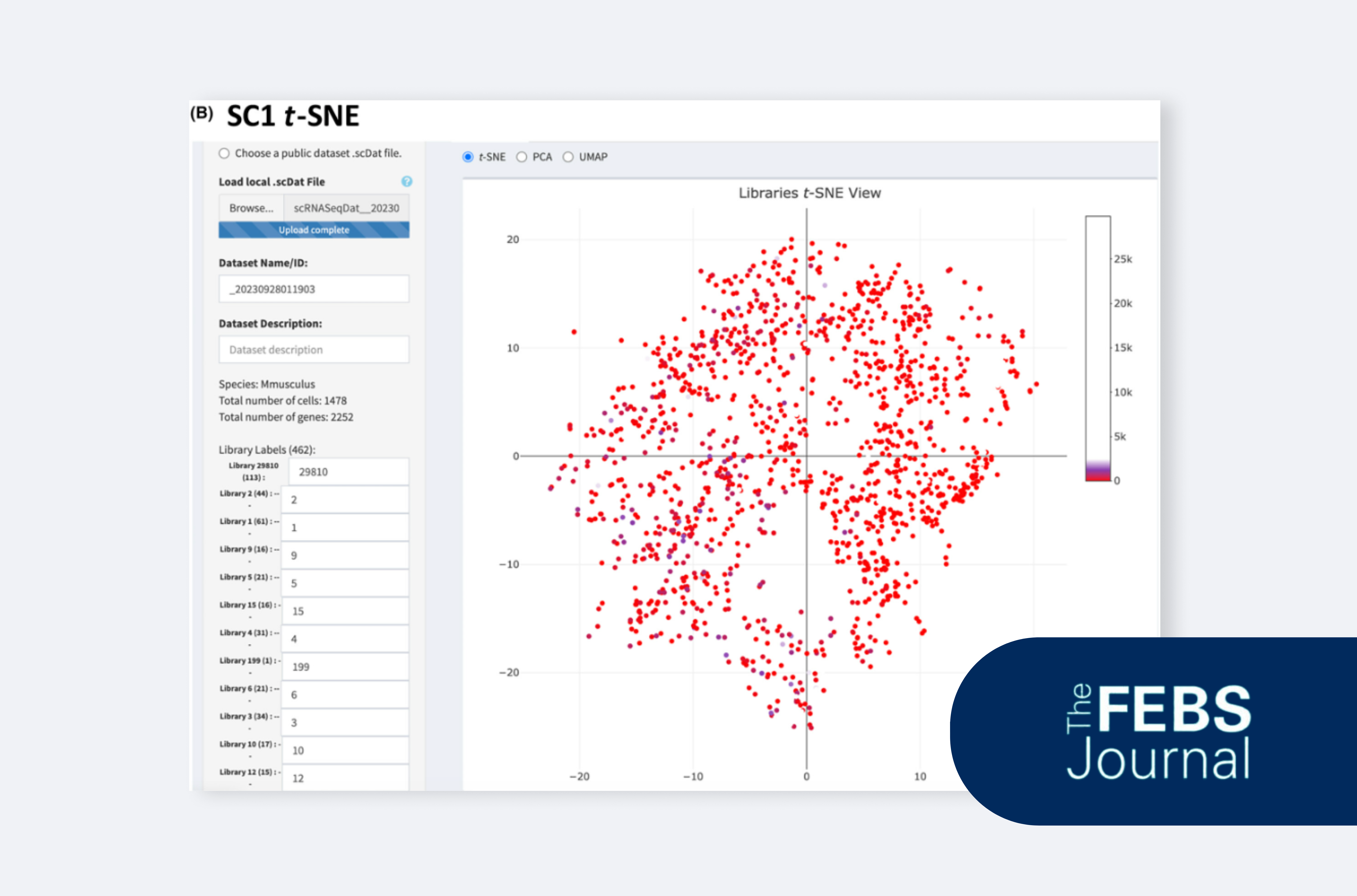
Sagnik Yarlagadda and Todd D. Giorgio provide a practical guide to scRNA-seq analysis using web-based platforms.
Read paper →