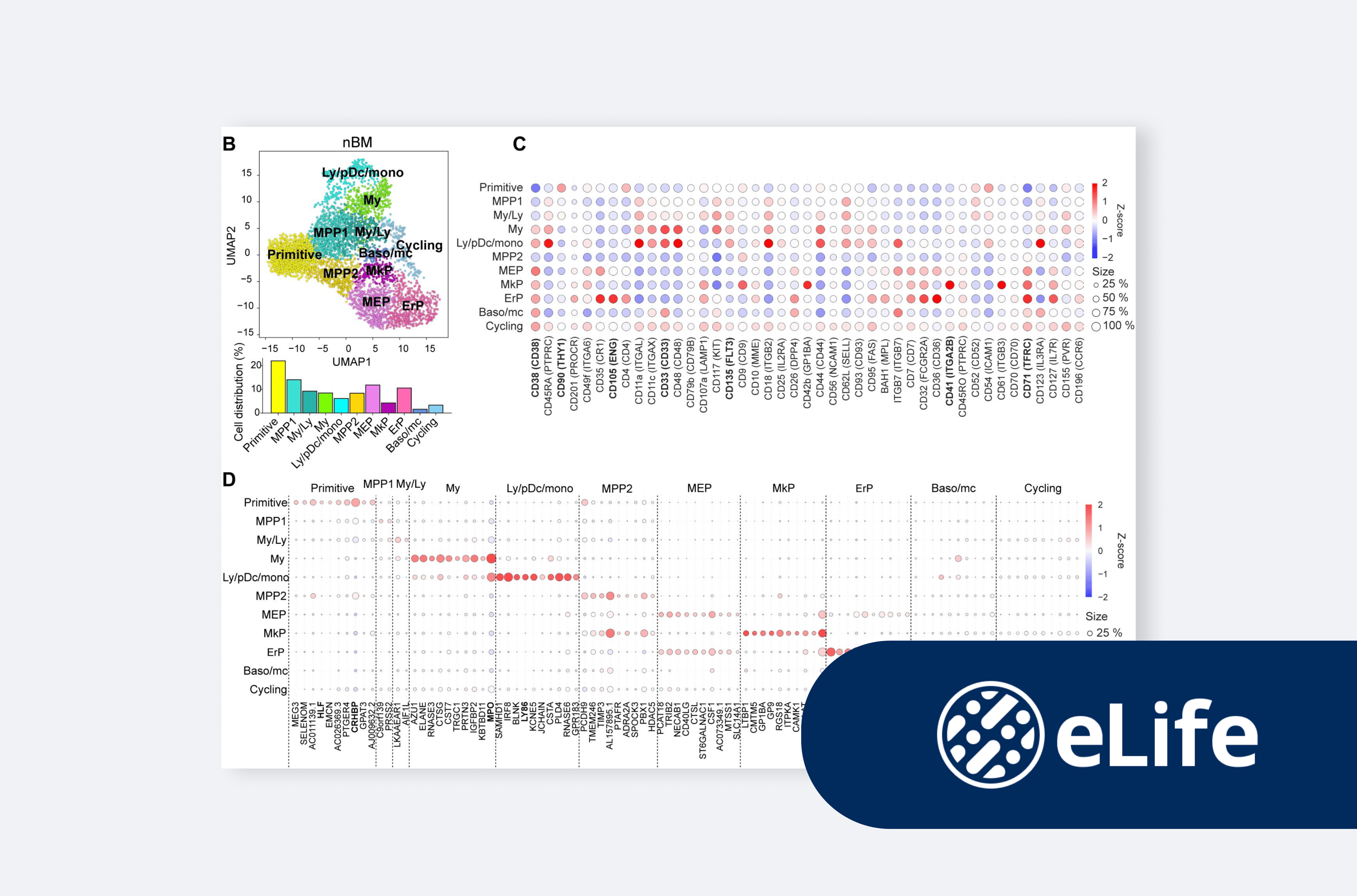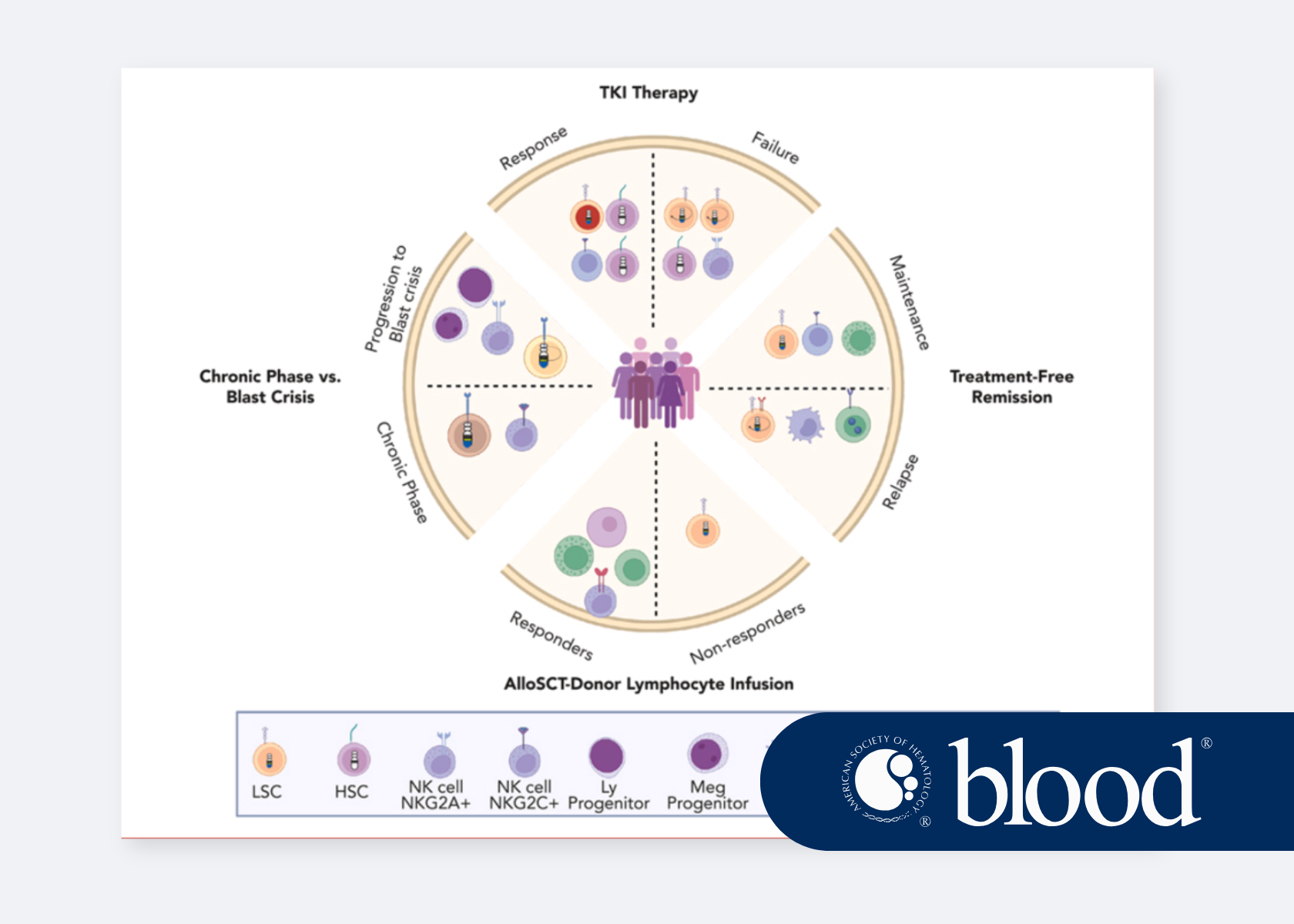
Insights from single-cell omics: cellular heterogeneity as a foundation of clinical outcome in chronic myeloid leukemia
Ram Krishna Thakur, Göran Karlsson explore how single-cell omics reveals cellular heterogeneity driving clinical outcomes in CML.
Read paper →