
Accelerated Single-Cell Data Analysis | Free Course by Nygen
A free, hands-on single-cell RNA-seq and multi-omics course. Learn to go from raw data to biologically interpretable results in under 3 hours using ScarfWeb.
View course →A curated index of articles, updates, events, and published work across single-cell research.

A free, hands-on single-cell RNA-seq and multi-omics course. Learn to go from raw data to biologically interpretable results in under 3 hours using ScarfWeb.
View course →
Deep learning and agentic AI analysis of single-cell RNAseq data.
Read more →
Learn how to identify marker genes in single-cell RNA-seq data without coding using Nygen's intuitive platform. This step-by-step guide covers data upload, quality control, clustering, marker detection, and dynamic analysis with detailed screenshots and expert tips.
Read more →
Uncover the essentials of marker gene identification in single-cell RNA sequencing. This article covers no-code solutions, methodologies, and case studies across various fields.
Read more →
Explore key challenges and advanced strategies in scRNA-seq data analysis for both new and experienced researchers.
Read more →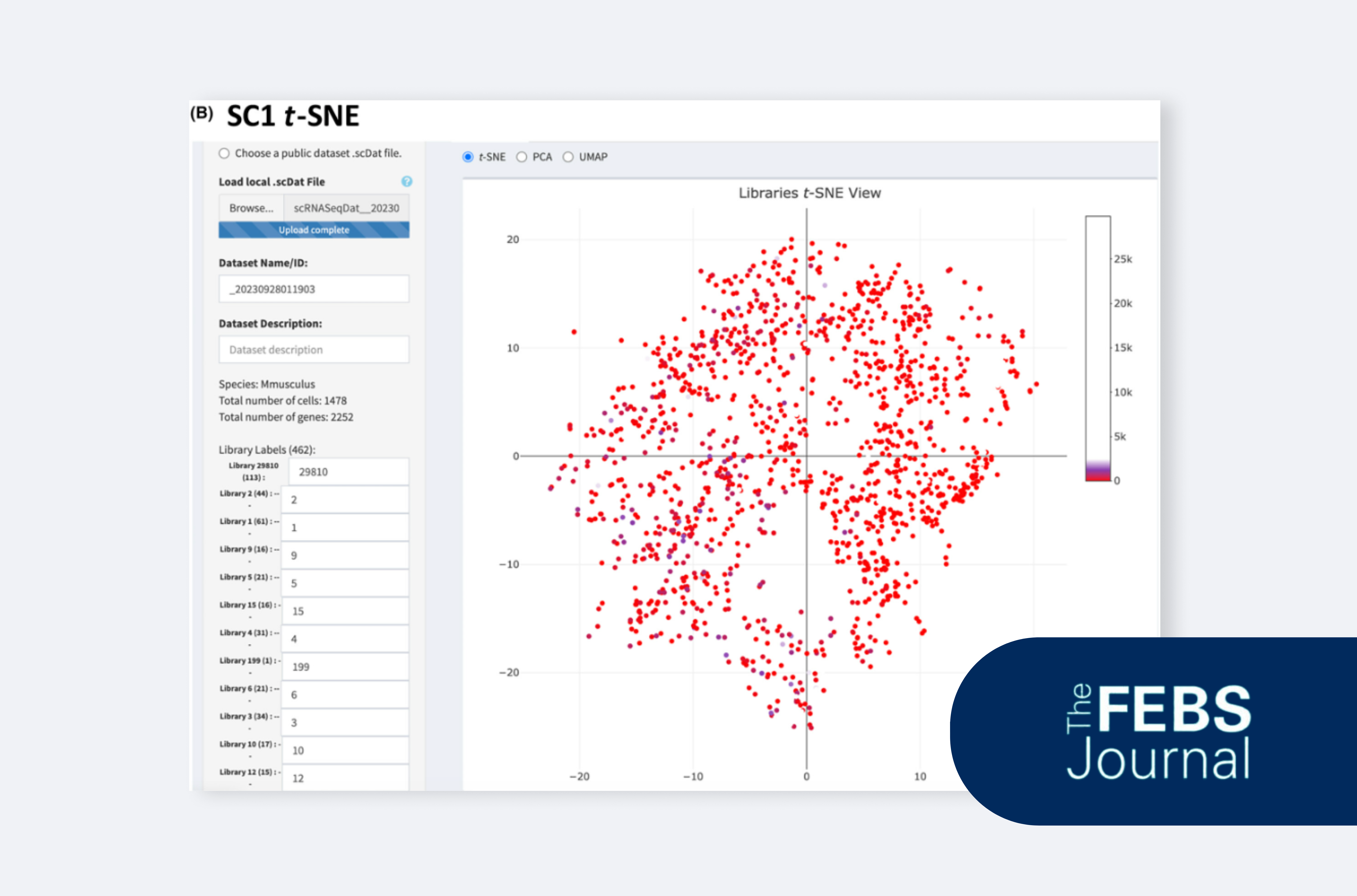
Sagnik Yarlagadda and Todd D. Giorgio provide a practical guide to scRNA-seq analysis using web-based platforms.
Read paper →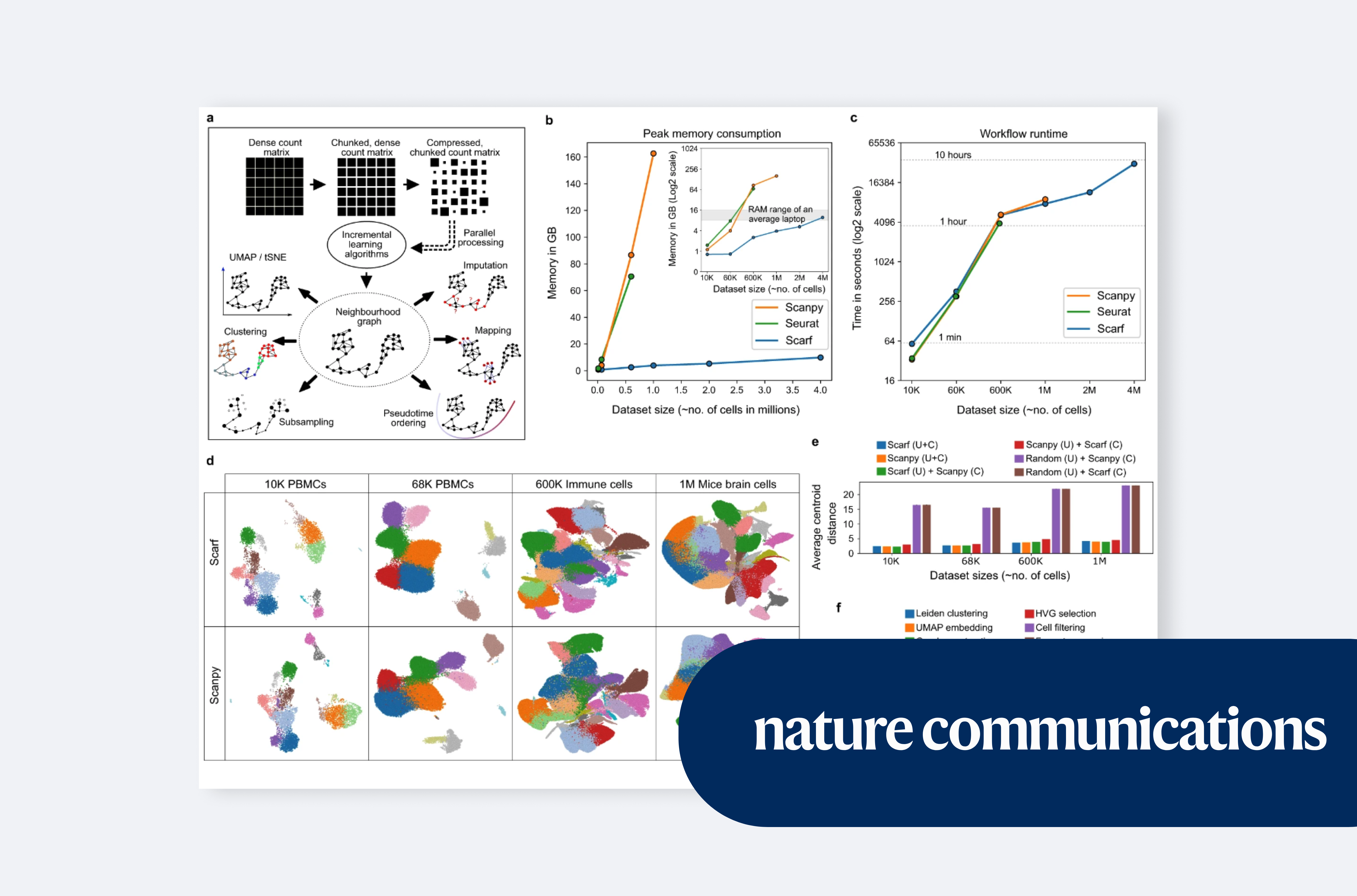
Parashar Dhapola et al. introduce Scarf for memory-efficient single-cell sequencing analysis, published in Nature Communications.
Read paper →