A Practical Guide to Single-Cell RNA-Seq Cluster Annotation
A practical guide to scRNA-seq cluster annotation in 2026, covering marker gene inspection, reference atlas mapping, supervised classification tools, and strategies for resolving ambiguous or disease-associated populations.
A Practical Guide to Single-Cell RNA-Seq Cluster Annotation
In this guide, we explore practical strategies for scRNA-seq cluster annotation, integrating marker gene sets, reference atlases, and supervised computational methods to assign reliable biological identities to individual cell populations.
Single-Cell RNA Sequencing (scRNA-seq): From Clustering to Meaningful Annotation
Single-cell RNA sequencing (scRNA-seq) has changed how researchers examine cellular heterogeneity by resolving gene expression at the level of individual cells. Clustering groups cells with similar transcriptional profiles, but clusters are only the starting point. The real biological question is what those clusters represent: established cell types, activation programs, transitional states, proliferative compartments, tissue-resident subpopulations, or disease-associated phenotypes.
Cluster annotation is the step that turns grouped expression profiles into interpretable biology. In practice, that means combining marker-gene inspection, reference mapping, tissue context, and careful validation. The field has moved well beyond purely manual labeling, but the most reliable workflows still keep biology in the loop. Automated tools can accelerate annotation and improve consistency, yet final labels should still be judged against marker expression, known lineage structure, sample context, and the possibility that some populations do not fit cleanly into existing references.

Foundational Concepts in scRNA-seq and Clustering
A typical scRNA-seq workflow begins with cell isolation, molecular barcoding, sequencing, and construction of a gene-by-cell count matrix. That matrix is then filtered and normalized so downstream analyses reflect biology more than technical noise. Dimensionality reduction methods such as PCA and UMAP help organize the data in lower-dimensional space, while graph-based clustering methods such as Leiden or Louvain identify transcriptionally similar groups of cells.
Those groups are not guaranteed to be cell types in the strict sense. A cluster may reflect cell cycle, stress, dissociation effects, mitochondrial content, donor-specific signal, treatment response, or a real biological lineage. Good annotation therefore starts before annotation itself. If quality control is weak, ambient RNA is not addressed, doublets remain in the data, or batch effects dominate the structure, later labels will inherit those errors.
It is also worth keeping resolution in mind. Under-clustering can merge biologically distinct populations, while over-clustering can split one coherent cell type into several artificial groups. Annotation works best when clustering has already been tuned against the biological question, the expected heterogeneity of the tissue, and the depth and quality of the dataset.
We explore these upstream analytical steps in more detail in our article on navigating scRNA-seq data analysis, and batch handling is covered further in the article on batch effect normalization techniques in scRNA-seq.

Why Annotation Matters
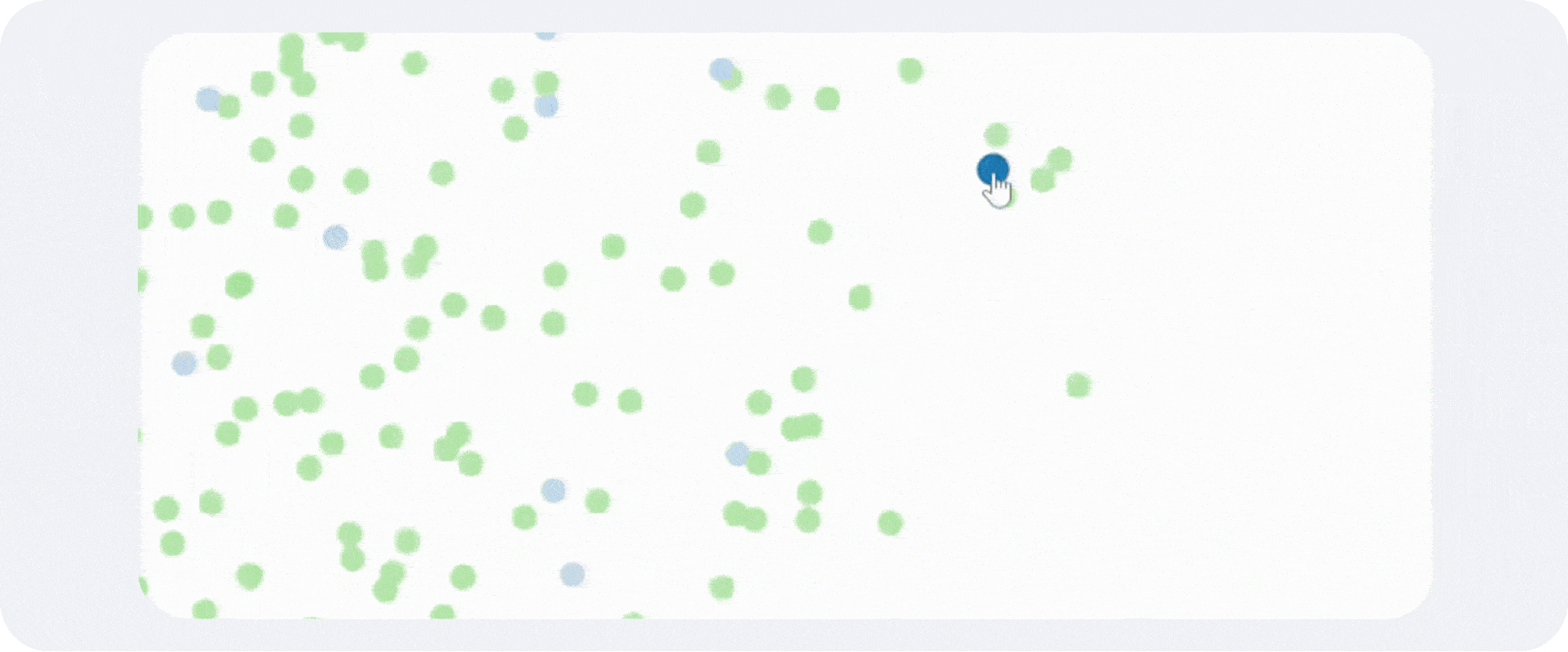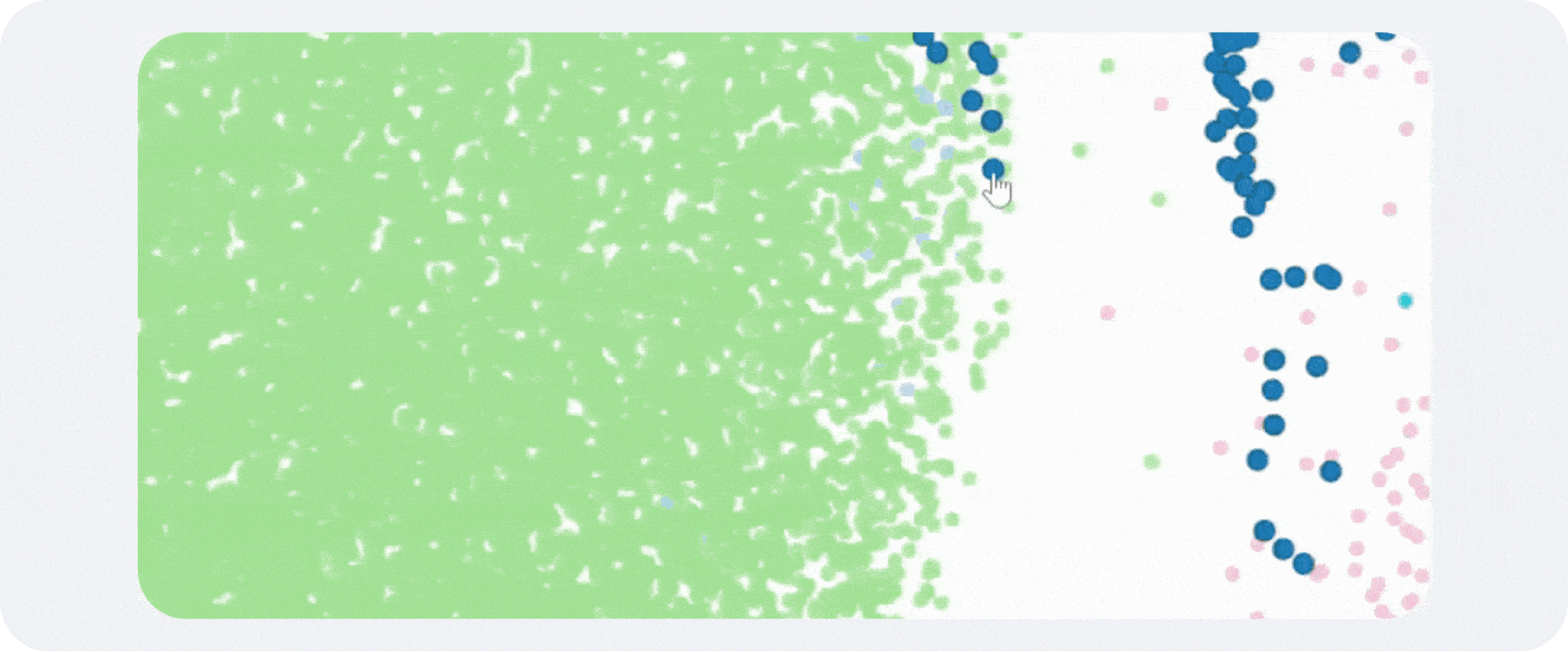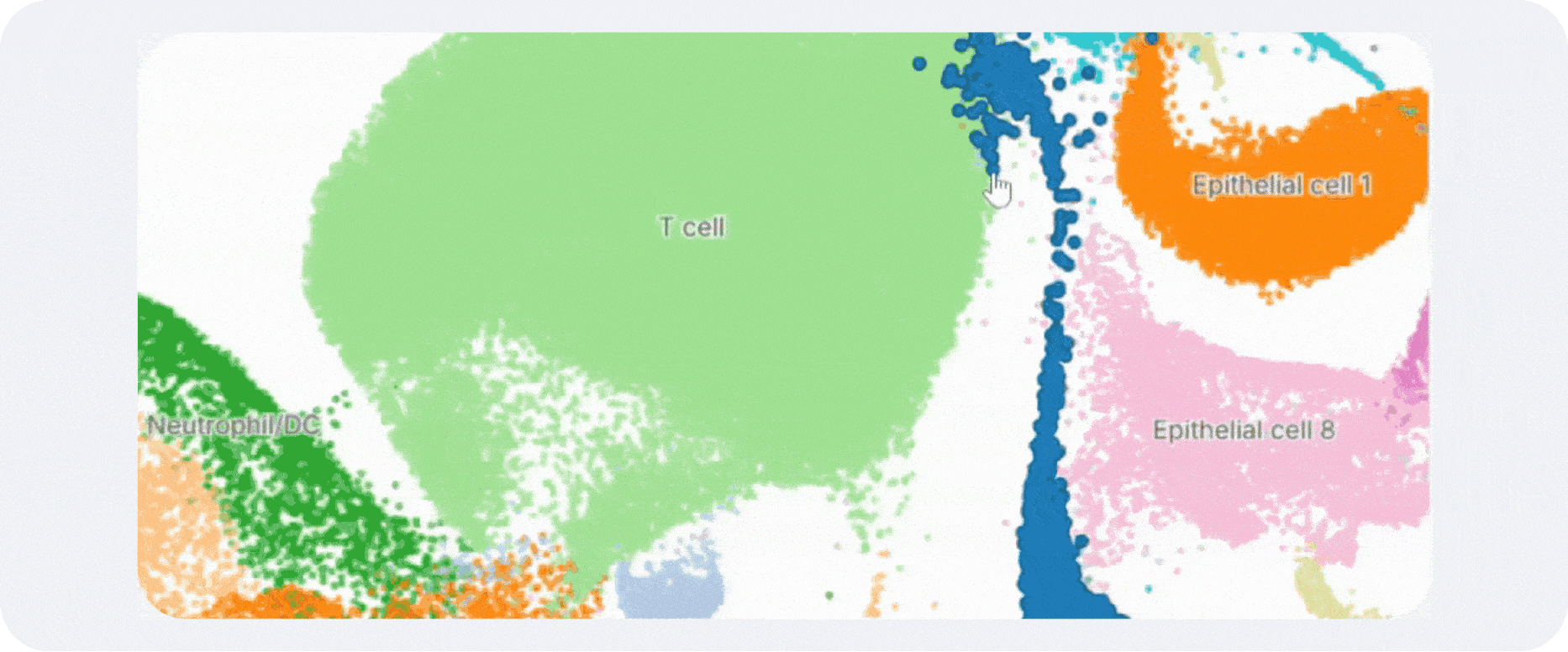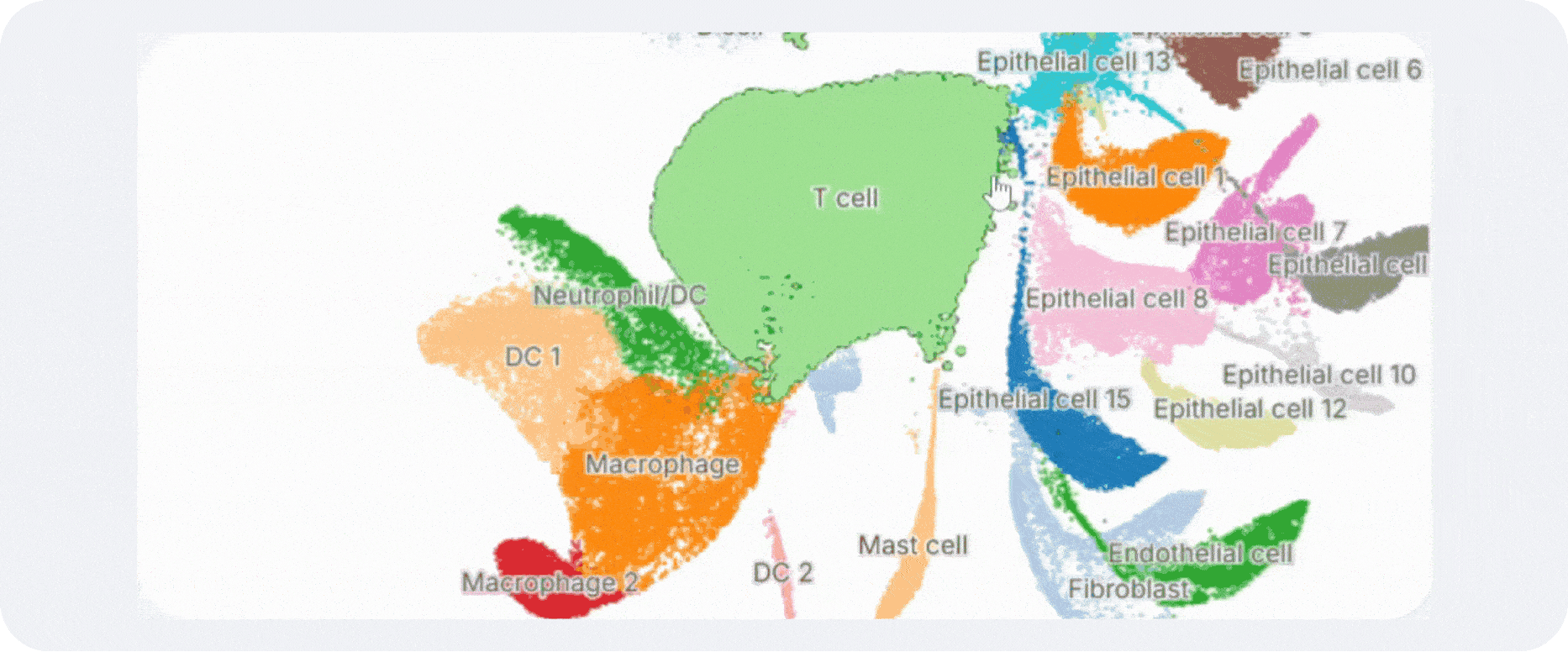
Annotation is what makes a clustering result usable. Without labels, clusters remain abstract coordinates on a UMAP. With good labels, they become epithelial subsets, activated T-cell states, stromal niches, developmental intermediates, malignant compartments, or whatever biology is actually present in the sample.
This matters for several reasons. First, annotation determines what downstream comparisons mean. Differential expression between two unnamed clusters is rarely useful on its own. Differential expression between inflammatory monocytes and tissue-resident macrophages is interpretable. Second, annotation affects reproducibility across studies. Shared naming and ontology-aware labels make it easier to compare datasets, integrate cohorts, and assess whether the same biology appears across tissues, conditions, or labs. Third, annotation quality often determines whether rare or clinically relevant populations are recognized at all.
In disease datasets, the stakes are higher. Tumor cells can occupy non-canonical states, immune cells can become context-specific, and damaged tissue can produce mixed or partial programs that do not match tidy textbook identities. In those settings, annotation is not only a labeling exercise. It is a biological interpretation problem.
Practical Strategies for Cluster Annotation
Begin with a biology-first perspective
A strong annotation workflow still begins with marker inspection. Start with cluster-level marker genes, then assess whether those markers support a coherent lineage and state. Canonical markers remain useful, but they should not be applied mechanically. Many markers are shared across related populations, and some become unreliable outside the tissue or disease context in which they were originally established.
Databases such as CellMarker 2.0 can help identify curated marker sets for human and mouse cell types. They are useful starting points, especially when entering an unfamiliar tissue, but they are references rather than truth. Marker interpretation should always be grounded in the sample type, known biology, and the combination of markers rather than a single gene.
In practice, ask a few basic questions: - Do the top markers support one lineage clearly? - Are there conflicting markers that suggest doublets, contamination, or transitional states? - Is the cluster defined by function, activation, proliferation, interferon response, stress, or tissue residency rather than a classical cell type? - Does the label reflect a broad class, a subtype, or a state, and is that level of specificity justified by the data?
Use curated marker-based and supervised tools
Once manual review establishes the broad structure of the dataset, supervised tools can accelerate the rest. SingleR compares query profiles against reference transcriptomes. CellTypist uses trained models for automated cell identity prediction. Garnett supports marker-based classification with user-defined marker hierarchies. These tools can be very effective when the reference is appropriate and the tissue context is represented well.
The important point is not to treat automated outputs as final labels. They are evidence. A good workflow asks whether the prediction matches the cluster markers, whether confidence is high across the cluster, whether nearby clusters receive similar competing labels, and whether the model is being forced to assign a known class to a genuinely unusual population.
Use reference atlases and label transfer carefully
Reference mapping has become a standard part of annotation. Resources such as the Human Cell Atlas and Azimuth make it easier to project query data onto well-characterized references and transfer labels based on transcriptional similarity. This can be especially helpful in common tissues where strong atlases already exist.
Still, label transfer has limits. A reference atlas may not contain the disease state, perturbation context, species, developmental stage, or technical chemistry represented in the query dataset. A transferred label can therefore be directionally helpful while still being biologically incomplete. It is often better to use the reference to anchor broad identity first, then refine subtype or state labels with local marker analysis.
Ontology-aware labeling is also increasingly important. The Cell Ontology provides standardized hierarchical terms for cell types, which helps make labels more consistent across datasets and publications. In practice, this often means separating the stable cell identity from the more contextual state. For example, a label such as "CD8-positive, alpha-beta T cell" may be the ontology-grounded identity, while "interferon-responsive" or "cycling" captures the local state layered on top.
Validate and iterate
Annotation is iterative. After an initial pass, revisit the cluster structure, marker genes, and nearest-neighbor relationships. Subset broad compartments when needed. Re-cluster lymphoid, myeloid, stromal, or malignant populations separately if the main embedding has compressed meaningful substructure. Check whether low-confidence labels become clearer after subsetting and whether broad labels fragment into interpretable subtypes.
Validation can also extend beyond transcriptomics. Flow cytometry, immunostaining, spatial context, perturbation metadata, lineage tracing, and orthogonal omics can all help resolve uncertain identities. That is especially relevant for transitional states and disease-associated populations, where transcriptional similarity alone is sometimes not enough.
Why Annotation Is Often Difficult
Below is a practical summary of common scRNA-seq cluster annotation challenges and how to handle them.
| Challenge | Description | What to watch for | Practical approaches |
|---|---|---|---|
| Batch effects | Technical differences between experiments, donors, runs, or chemistries can create artificial separation or hide real structure. | Labels that align more with sample origin than biology. | Apply careful QC and integration strategies such as Harmony, MNN-based correction, or Seurat integration, then re-check marker coherence. |
| Ambiguous marker genes | Some populations lack unique markers, and many closely related cell types share canonical genes. | A cluster appears plausible for several related identities. | Use combinations of markers, pathway context, and neighborhood structure rather than relying on one or two genes. Cross-check with curated databases and literature. |
| Rare cell populations | Low-abundance populations may be masked by dominant lineages or lost through conservative filtering. | Small clusters with distinct markers are easy to dismiss as noise. | Test multiple clustering resolutions, inspect rare-marker expression carefully, and validate with orthogonal methods where possible. |
| Transitional states | Differentiating or activated cells may express markers from more than one lineage. | Mixed signatures that do not fit a stable endpoint identity. | Use trajectory-aware tools such as Monocle, Slingshot, or PAGA and describe the cluster as a state when a fixed cell type is not justified. |
| Disease context | Cancer, inflammation, infection, and injury can produce non-canonical transcriptional programs. | Reference-based labels look close but incomplete or misleading. | Use disease-relevant references when available, incorporate pathway analysis, and separate stable identity from disease-associated state. |
| Biological vs technical variation | Stress, dissociation artifacts, ambient RNA, and doublets can mimic real populations. | Clusters dominated by ribosomal, mitochondrial, hemoglobin, or mixed-lineage signatures. | Revisit QC, ambient RNA correction, and doublet detection before forcing a biological label. |
| Cross-species differences | Human references are often applied to non-human datasets with uneven success. | Conserved broad lineages but unstable subtype assignments. | Use species-matched references where possible and be conservative with fine-grained labels. |
| Incomplete references | Existing atlases do not capture every tissue, perturbation, age, or disease setting. | Automated tools assign the nearest known class even when the fit is weak. | Treat the prediction as a clue, not a conclusion. Document uncertainty and retain descriptive labels when appropriate. |
| Overconfident automation | Models can assign precise labels even when the evidence is weak. | Clean-looking labels with poor marker support. | Require agreement between model output, markers, and tissue context. Keep uncertain populations explicitly marked as such. |
| Interpretation overload | Large datasets with many clusters can become hard to review systematically. | Inconsistent annotation depth and undocumented decisions. | Use structured review workflows, marker panels, ontology mapping, and annotation notes so decisions remain traceable. |
Future Directions and Innovations
The annotation landscape is moving in a few clear directions.
Multi-omic and spatial context
Annotation is increasingly informed by more than RNA alone. CITE-seq, ATAC-RNA multiome data, and spatial transcriptomics can help distinguish populations that look similar in transcript space but diverge in protein expression, chromatin accessibility, localization, or tissue architecture. This is especially useful for immune activation states, stromal heterogeneity, and tumor ecosystems where transcription alone can flatten meaningful differences.
Better reference resources
Reference atlases continue to improve in both size and tissue coverage. The Human Cell Atlas remains a major organizing effort, and tools like Azimuth have made reference mapping more practical in everyday analysis workflows. As these references expand into diseased, developmental, and perturbational settings, automated annotation should become more informative, though not necessarily less dependent on expert review.
More structured annotation outputs
There is a gradual shift away from single flat labels toward richer annotation objects: broad lineage, finer subtype, cell state, confidence level, and evidence trail. That structure is useful because it reflects how annotation actually works. A cluster can be confidently myeloid, probably monocyte-derived, and only tentatively inflammatory.
AI-assisted interpretation, with caution
AI methods are becoming more visible in annotation workflows, particularly for literature-supported interpretation, marker synthesis, and assisted review. The useful role for these systems is not to replace biological judgment but to reduce manual overhead, surface relevant evidence faster, and help standardize reasoning across large studies. The hard problem remains the same: tying predicted labels to defensible biological evidence rather than producing polished but weakly grounded assignments.
Best Practices and Recommendations
A practical annotation workflow in 2026 usually follows a pattern like this:
- Start with data quality. Remove low-quality cells, likely doublets, and obvious technical artifacts before treating any cluster as biological.
- Inspect markers manually. Build an initial view of major lineages and obvious cell states from cluster markers.
- Run supervised tools and reference mapping. Use SingleR, CellTypist, Azimuth, or similar methods to generate supporting evidence, not final truth.
- Separate identity from state. Distinguish core cell type from activation, cycling, interferon response, stress, or disease-associated programs.
- Use ontology-aware labels where possible. Standard terms improve reproducibility and downstream comparison.
- Subset and re-cluster when needed. Broad compartments often contain interpretable heterogeneity that is hidden in the full embedding.
- Document uncertainty. It is better to leave a label at "fibroblast-like stromal cell" or "cycling T/NK population" than to overstate precision.
- Validate against orthogonal evidence. Where the biology matters, support labels with protein, spatial, lineage, or perturbation data.
Looking Ahead: Can AI Evolve Beyond Statistical Prediction?
Current annotation methods are still strongest when they combine statistical matching with biological review. That has not changed. What has changed is the amount of evidence that can now be brought into the loop. References are larger, ontologies are more usable, supervised tools are more accessible, and AI-assisted systems can help researchers move faster through literature, marker interpretation, and cross-dataset comparison.
But the central challenge remains biological, not cosmetic. A good annotation is not the most specific label a model can produce. It is the most defensible label the data can support.
For teams working at scale, the practical goal is not full automation. It is a workflow where evidence from markers, references, atlases, and computational models can be reviewed in one place and turned into labels that are both biologically honest and analytically useful.



